Introduction
Numerous studies have reported the association between intestinal dysbiosis and the development of cancer [1–8] and the importance of the microbiome in cancer development [9,10]. Intestinal dysbiosis is defined as an alteration in the composition and/or richness (i.e., the number of unique bacterial species) of the intestinal microbiota [11]. It has been proposed that the microbiome is involved in the initiation and progression of various types of cancer [12] and that the tumor and tumor site themselves have a distinct microbiome in certain cancers [13]. In addition, certain microbial agents or their products have been shown to have anticancer effects [14,15]. Modulation of the microbiome has been shown to be helpful in the treatment of certain types of cancer [15]. Despite all of the evidence linking the intestinal microbiome with cancer, there remains a paucity of studies evaluating the effects of the microbiome and its product on the development or treatment of metastasis.
From its primary location to distant parts of the body, the spread of cancer is the most common cause of mortality for solid tumor cancer patients [16]. Progress has been made to clarify that discrete steps are taken by cancer cells within a complex set of events (i.e., the metastatic cascade) involving interactions between tumor cells, normal stromal cells recruited to the tumor microenvironment, and cells of the immune system. This complex sequence of events is coordinated and triggered remains unclear; nonetheless, the complexity of these interactions is unlikely to be explained by tumor cell determinants alone and more likely from the interaction between tumor cells, the tumor microenvironment, and the host [17]. Here, we now ask if the host contribution to trigger and regulate these cellular interactions may involve the gut microbiota. As we propose this new research agenda, we hypothesize that modulation of the metastasis microenvironment resulting from gut microbiome alterations influences the metastatic cascade.
Given the growing understanding of the regulatory influence of the gut microbiome on both stromal and immune cells during wound healing [18,19], embryogenesis, and inflammatory responses [20–25], we now ask if alterations in the gut microbiome may be a trigger for the cell-to-cell interactions associated with discrete steps in the metastatic cascade. Indeed, there is mounting evidence that the trillions of resident gut microbiota in the human gastrointestinal tract play a pivotal role in shaping host immune responses that may target micrometastatic cells, modulating stem cell localization, and inducing the release of metabolic signals. These three factors may individually or collectively modulate tumor metastasis. Thus, we believe that asking questions about the potential connection between the gut microbiome and cancer metastases is timely, needed, and supported by our understanding of both the cell biology of metastasis and the homeostasis of the gut microbiome. In this perspective piece, we will focus on three specific questions with regards to how alterations in the gut microbiota may influence cellular interactions linked to cancer metastasis:
- Could gut microbiota-derived signals or cues trigger the break-in metastatic dormancy?
- Are there patient-specific features regulating the pre-metastatic niche that are modulated by gut microbiota?
- Could the gut microbiome interaction with tumor cells through epigenomic/epigenetic regulation help predict metastatic proclivity?
Current Thoughts on Cancer Metastasis: Metastasis describes both the process of cancer spread (i.e., the verb) and the resultant secondary cancer lesion (i.e., noun). The process of metastasis involves microscopic tumor cells leaving the primary tumor. Through various proposed mechanisms, cells pass through the tumor matrix and then through or between endothelial cells to enter the circulation. While in the circulation, tumor cells must resist anoikis (programmed cell death associated with loss of cellular contact), evade immune recognition, demonstrate metastatic endurance by coping with and adapting to a variety of cellular stresses, and either immediately proliferating or entering dormancy, and then breaking dormancy to proliferate within the distant organ subsequently. The lack of mechanistic clarity around triggers that break dormancy and the formation of the pre-metastatic niche seems to be the most obvious link to a potential role of gut microbiota in modulating host immune responses and, ultimately, the development of tumor metastasis [19].
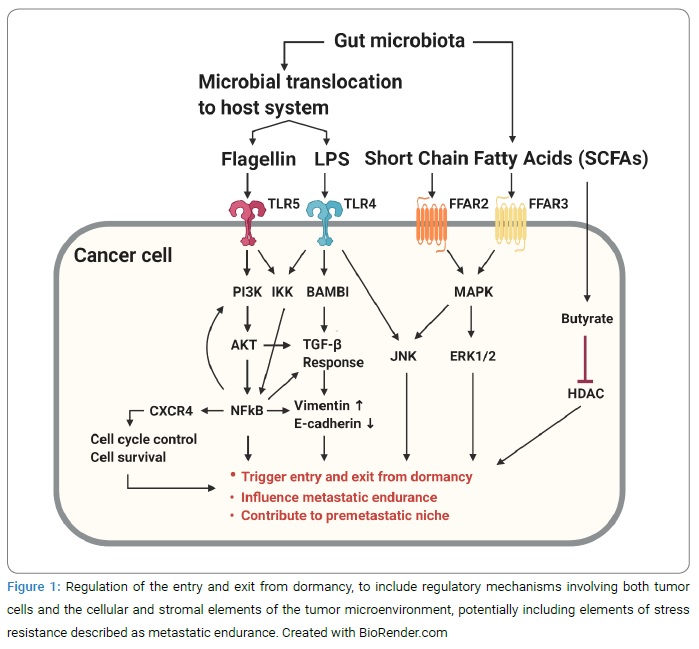
Metastatic Dormancy: Clinical observation in several solid tumors includes late metastatic recurrence of cancer despite complete resection of a primary tumor and often following adjuvant chemotherapy. This clinical scenario has been explained by micrometastatic tumor cells, leaving the primary tumor, entering a relatively chemo-resistant state associated with dormancy at distant organ sites or sanctuary sites (i.e., bone marrow), and then exiting this dormant state following an unknown trigger to proliferate and yield late metastatic recurrent lesions. It remains unclear how the entry and exit from dormancy are regulated but postulated to include regulatory mechanisms involving both tumor cells and the cellular and stromal elements of the tumor microenvironment, potentially including elements of stress resistance described as metastatic endurance (Figure 1, Figure 2).

Impact of gut microbiota on stem cell traits: Based on the existing understanding of the roles of the gut microbiota inducing or mobilizing stem cells in the setting of wound healing [18,19] and embryogenesis, it is reasonable to ask if similar events may serve as triggers or signals that may break the dormancy of micrometastatic cells.
For example, potential mechanisms that may be explored further include the impact of STAT and JNK pathway modulation resulting from gut microbiome alterations and impacts on metastasis biology (Figure 2). In studies in Drosophila, JAK-STAT and JNK pathways are required for bacteria-induced stem cell proliferation [26]. In fact, intestinal Pseudomonas aeruginosa infection in Ras1 oncogene expressing Drosophila causes a dramatic over-proliferation of intestinal stem cells and initiates ISC-mediated cell mobilization and survival through activation of the JNK-pathway [27].
Similarly, in mice, pathogenic bacteria such as Helicobacter pylori, Salmonella spp., and Citrobacter rodentium can induce JAK-STAT pathways [28–32]. Similarly, the commensal gut microbiota Bacteroides fragilis can induce β-catenin to localize to the nucleus, which results in the expression of genes required for stem cell traits that may be associated with the metastatic phenotype [33]. In addition, it has been recently reported that the microbiota-derived lactate stimulates ISC proliferation through Wnt/β-catenin signals of Paneth cells and intestinal stromal cells [34]. On the other hand, gut microbiota-produced butyrate inhibits stem cell proliferation in a transcription factor Foxo3 dependent manner [35]. These observations provide the basis for hypotheses that the gut microbiota may influence a cellular phenotype observed in metastatic dormancy and metastatic endurance by promoting the activation of the Wnt/β-catenin pathway. There are two possible mechanisms by which the gut microbiota could have these distant effects. The first is by contributing to the development of a specific, distinct tumor microbiome [36]. The second is via gut microbiota-derived metabolites.
Pre-metastatic Niche: In addition to the concept of dormancy over the past decade, several new research areas in the field of metastasis have emerged. These new insights have been added to Stephen Paget’s 1889 explanation for the non-random preference certain cancers have for growth at certain secondary sites [37]. Briefly, Paget’s seed and soil hypothesis suggested that the sites where metastases occur are defined not only by the tumor cell (“seed”) but also the microenvironment of the secondary metastatic site (“soil”). Recent data has shed new light on the interaction between the seed and soil. Interestingly, the formation of the friendly metastatic tumor microenvironment (i.e., the soil) can precede the arrival of metastatic cells by mobilizing bone marrow-derived cells to form the so-called premetastatic niche at the secondary site. It is interesting to consider that recruitment of cells forming the premetastatic niche may be coordinated through alterations in the gut microbiota in similar ways to those observed with the gut microbiome regulating wound healing and stem cell trafficking.
Metastatic Endurance: Upon arrival at a distant site, the new microenvironment encountered by the metastatic cell is considered foreign and hostile. Survival of metastatic cells at the secondary site is likely a consequence of intrinsic features of the metastatic cell and an ability to engage in a molecular cross-talk with its surroundings effectively and, in so doing, modulate the environment of the secondary site to allow cell survival in the setting of stress [38]. This engagement can occur by several mechanisms that we have collectively coined as metastatic endurance, which is also linked to the process of metastatic dormancy. Could the microbiome, through its effects on the immune system and metabolite production, influence the pathways that lead to the survival of metastatic cells?
Effects of microbiota on chronic systemic inflammation and link to cancer metastasis: The microbiota is involved in limiting systemic inflammation in the healthy state and contributing to systemic inflammation when intestinal dysbiosis is present [39,40]. Chronic systemic inflammation has been shown to lead to the development of many diseases, including cancer development and progression [41–43]. Distinct of this, we now propose a rationale to consider the association between intestinal dysbiosis and cancer metastasis biology. Multiple tumor types have been shown that high serum C-Reactive Protein (CRP) and inflammatory cytokines increase the risk of recurrence and metastasis [44,45]. Further evaluation of inflammation markers may help answer some of the unknown questions surrounding intestinal dysbiosis and its connection to cancer metastasis in general and some of the more novel and less understood components of the metastatic cascade.
Provocative Question 1: Could gut microbiota-derived signals or cues trigger the break-in metastatic dormancy?
Could there be an endocrine mechanism associated with the microbiota that takes cells out of dormancy or modulate non-cellular matrices (fibrinolysis, collagen type shifting?), similar to the observed effects of gut microbiota alterations during embryogenesis?
Provocative Question 2: Are there patient-specific features regulating the premetastatic niche, and are these modulated by gut microbiota?
The microbiota may be a trigger that affects immune surveillance in the setting of metastasis. The microbiota has a significant role in modulating the immune system. One way in which this occurs is their role in increasing regulatory T cells (Tregs). Normal commensal bacteria in the gut are necessary for the production of Tregs [46,47]. Tregs act to suppress the immune response and maintain immune homeostasis [48,49]. The number of Tregs is increased in nearly all cancers in order to damp responses from the immune system against cancer cells, metastasis, tumor recurrence, and treatment resistance [49].
Further research is necessary to look at the direct mechanisms that implicate intestinal dysbiosis as a contributing factor to the dysregulation of the local immune response during metastasis. Indeed, metastatic inefficiency may be the result of immune rejection of metastatic cells or local immune suppression at discrete secondary sites. The generation of local immunosuppression as part of the development of the premetastatic niche may produce pro-inflammatory cytokines from the progressive metastatic burden that could further dysregulate the intestinal microbiome. This could become a vicious cycle leading to further progression of metastasis. Could gut dysbiosis be characterized by a predominance of gut bacteria known to induce stem cell traits (i.e., Escherichia coli, Pseudomonas aeruginosa, Helicobacter pylori, and Salmonella spp.) promote metastasis by altering the pre-metastatic niche? This begs the question as to whether monitoring these or other specific gut microbiota could be used as potential biomarkers in identifying patients with a higher risk of metastasis.
Provocative Question 3: Could the gut microbiome interaction with tumor cells through epigenomic/epigenetic regulation help predict metastatic proclivity?
The metastatic proclivity of tumor cells is determined by both tumor cell-intrinsic factors and extrinsic factors from the tumor microenvironment. Even though tumor cells’ intrinsic properties have been considered the driving force for metastatic processes, the cancer genotype often struggles to predict metastatic proclivity alone. Non-genomic events (including epigenomics) may provide clarity on the metastatic program of cancer cells [50,51]. Given the role of gut bacteria on mechanisms of epigenomic controls, we now ask if triggers for epigenomic modifications of metastasis-associated genes are directed by changes in the gut microbiome, inclusive of the recognized role of specific gut microbiota-derived metabolites on modulation of the complex histone assembly.
Gut microbiota-derived metabolites and possible linkage to metastasis: Are there certain metabolites produced by the gut microbiota that could be contributing to the break from metastatic dormancy, development of the premetastatic niche, or epigenetic regulation of the metastatic tumor cells? (Figure 3) The basis of this hypothesis is found in the role that bacterial metabolites play in oncogenesis and their role in post-transcriptional regulation.
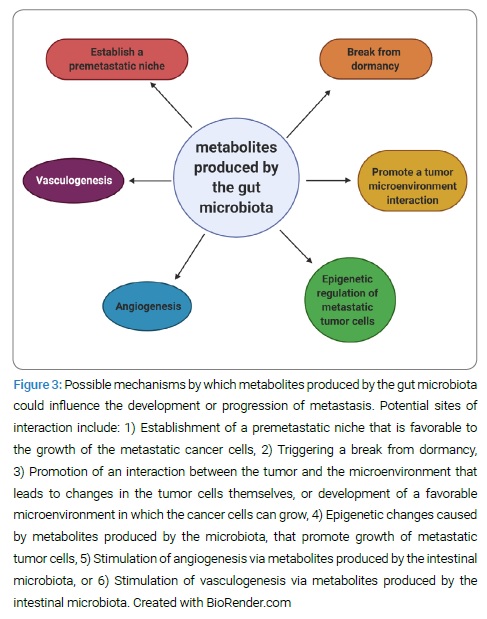
Bacterial metabolism and metabolite production appear to be crucial factors that serve as a foundation for the observed associations between diet and various cancers [52]. In colorectal cancer, gut bacterial fermentation of dietary fiber to Short-Chain Fatty Acids (SCFA), such as butyrate, is believed to play an important role in suppressing oncogenesis via its anti-inflammatory and antiproliferative effects [53]. In contrast, bacterial metabolism of bile acids and proteins can result in carcinogenic aromatic amines and sulfides [52]. Further studies are warranted to evaluate the effects of these and other metabolites in the setting of metastasis.
Intestinal dysbiosis has been shown to operate through intermediaries to alter the health of distant organs. An example is the production of Short-Chain Fatty Acids (SCFAs). SCFA can act as inhibitors of histone deacetylases to induce hyperacetylation of histones which affects gene expression and results in anti-inflammatory properties, induction of growth arrest, and apoptosis [54]. Histone Deacetylase (HDAC) inhibitors induce cancer cell cycle arrest, promote differentiation and cell death, reduce angiogenesis, modulate the immune response, and induce apoptosis of tumor cells, with minimal effects on normal tissue [55,56]. In addition to decompaction of the histone/DNA complex, HDAC inhibition also affects the acetylation status and function of non-histone proteins. HDAC inhibitor drugs have demonstrated antitumor activity in clinical trials [55]. It has been shown that gut microbiota-produced butyrate directly binds to HDAC and inhibits HDAC activity, which significantly affects the epigenetic regulation of host gene expression [77]. It has been proposed that the effects of specialized metabolites may explain current knowledge gaps in linking the gut microbiota to biological host mechanisms in various disease states, including cancer. We now propose that these or other metabolites may influence metastasis progression [57,58].
The questions that will need to be addressed in future studies include:
- What are the cellular phenotypes involved, and how are they regulated?
- What are the bioactive metabolites produced in dysbiosis, and what are their targets?
- Are there metabolites from the microbiota involved in establishing a premetastatic niche, triggering the end of dormancy, or promoting a tumor microenvironment interaction?
- Are there secreted bioactive metabolites that contribute to vasculogenesis, angiogenesis, or epigenetic regulation of metastatic tumor cells?
Proposal for development of a novel dysbiosis blood test: The intestinal microbiome acts as a functioning organ with a variety of interactions with the mammalian body. Through analysis of metabolites and other blood biomarkers, we will continue to gain insight into the microbiome’s function and its effects on health and disease [59–63]. In addition to identifying the specific bacteria that contribute to carcinogenesis, identifying the mediators through which these bacteria promote cancer is essential to advance therapeutic interventions. If there are signals secreted by the gut microbiota that are influencing metastasis, what are they? Potential candidates include proteins, microRNA, or certain metabolites. Can the measurement of metabolites produced by gut microbiota, such as SCFAs, indole derivatives, bile acid metabolites, choline metabolites, phenolic derivatives, and polyamines, help us distinguish between a normal state and intestinal dysbiosis? If an association with certain metabolites is found, could this be a potential link between intestinal dysbiosis and the development of cancer or metastasis? For example, Indole-3-Propionic Acid (IPA) is a powerful antioxidant that exhibits cytoprotective effects. Could low levels of IPA contribute to cancer growth or metastasis? Certain vitamins (B1, B2, B3, B5, B6, B7, B9, B11, B12, K2) are important enzymatic cofactors for DNA replication, repair, methylation, and regulation of cell proliferation [64]. We suggest that there may be a metabolite or set of metabolites that are produced in the GI tract, absorbed into the systemic circulation, and affecting the tumor cells’ ability to metastasize, either directly or through other host cells (Figure 1, Figure 2). Alternatively, or in addition to this, there may be immune system markers (cytokines, Treg cells, etc.) that will help us identify if and how the intestinal microbiome is affecting cancer metastasis.
Possible future therapeutic applications: The mechanisms by which the microbiota may influence metastasis and the break-in dormancy may uncover therapeutic and prognostic tools useful in improving outcomes for cancer patients. Although additional data linking the microbiome’s contribution to specific cancers and metastasis of cancer need to be generated, microbiota-based therapeutic strategies for cancer prevention and treatment have been proposed, and some are already in use [6,8,65–67]. Dietary intervention, prebiotics, probiotics, or fecal microbiota transplant may restore gut microbiota homeostasis and thus influence various physiologic phenotypes. Understanding the diverse contributions of the bacterial microbiota to metastasis will open new possibilities for diagnostic, preventative, and therapeutic approaches.
Microbiota can improve cancer therapeutic responses [68,69]. The influence of the gut microbiota in shaping local and systemic immune responses has been recognized [70]. The effect of this biological function of the microbiota on the efficacy of antitumor agents has begun to be studied. The effect of gut microbiota on the efficacy of chemotherapy and immune checkpoint inhibitor therapy has been investigated, and results have suggested that the gut microbiota are an important covariate of treatment response [20,24,25,71]. Optimizing the function of the gut microbiota may improve or augment existing cancer therapy and maybe a missing link in explaining the seemingly currently unpredictable responses observed in the field of immunotherapy.
How comparative oncology in the dog offers potential answers: Studies to understand and then demonstrate a connection between gut microbiome alterations and cancer metastasis will lead to the need to demonstrate a clinical value to cancer patients and patients with cancer metastasis. The many questions that need to be addressed within this research agenda cannot be answered in human clinical trials or conventional preclinical studies alone; new approaches are needed. The naturally occurring metastatic cancers in pet dogs may allow these important questions to be answered. There are many physiologic similarities between canine cancer and metastasis and human cancer and metastasis [72,73]. There are also many similarities between the canine and human microbiota. In order to evaluate the effects of the microbiome on metastasis, it is reasonable to consider the dog as a model for human disease. Canine models for human microbiome alterations are promising due to their similarity in environmental conditions, diet macronutrients, genetic heterogeneity, microbiota composition, and function. Murine models are widely used for assessing changes in the human microbiome [74]; however, they do not accurately mimic the human condition due to genetic homogeneity and differences in gut microbiome composition. The canine model offers greater genetic heterogeneity than murine models. This heterogeneity will be more representative and reflective of human disease when compared to studies in mice. The dog microbiome is closest to the human microbiome, based on a study that compared dogs, humans, pigs, and mice [75]. Findings in future dog studies may therefore be predictive of human microbiome results. The canine has similarities in gut morphology and function compared to humans’ digestive system, which lends itself to the potential of a good disease model. Perhaps more important than the organisms present, the metabolic capacity of gut microbial communities of canines, humans, and mice shows that the canine microbiome function is more closely related to the human than the mouse microbiome [76].
An additional value in using dogs as a model for human disease in metastasis is the shorter time frame in which metastasis occurs in dogs in many cancer models compared to humans. The questions that we would like to ask in the setting of human metastasis can be asked in pet dogs in ways that are not feasible in humans or conventional preclinical models alone. Studies in dogs could supplement other preclinical data and bolster clinical investigations in tumor types for which there is a paucity of human patients for clinical trials.
Future studies will be required to provide a deeper understanding and characterization of the metagenome and metabolome of dogs in healthy and diseased states. After additional baseline data is established in health and disease, we will then evaluate the effects of specific dietary and other therapeutic interventions.



